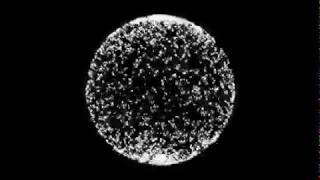
Actual things start happening around the 1:20 mark. Note that concentric waves are generated first, which later turn into spiral waves as cells start to lose the ability to oscilate on their own. The video is sped up 100 times.
this simulation of cAMP waves made by reimplementing the model described by Lauzeral et al. in "Desynchronization of cells on the developmental path triggers the formation of spiral waves of cAMP during Dictyostelium aggregation", published in the proceedings of the natural acadamy of sciences USA, volume 94, pages 9153-9158, August 1997
The model was implemented in C++, using openCV to visualize the data. 1280x720 voxels were simulated, each representing 0.1mm by 0.1mm. The timestep was 1 second per iteration. The desynchronization parameter in this run is 25 minutes.
Code: [ Ссылка ]
To compile, first set up include paths and linker settings for openCV. I think this was with openCV 3.0.0 beta, but any changes in openCV should lead to only minor required corrections in the code.