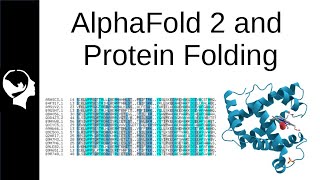
Deepmind have recently cracked the more than 90% GDT on protein folding estimation. Traditionally protein structured are determined experimentally through X ray Crystallography, which takes approximately $120,000 and almost a year to complete, additionally, many proteins are not able to be crystallized and unable to be studied this way.
Alpha fold 2 used a mixture of attention based neural network (transformers), multiple sequence alignment, and amino acid residue pairs for the model building. The model is then train from end to end on 170,000 3D protein structures and more than 17 millions protein sequences to establish the final output.
Estimation of protein structure can have significant impact on the Proteinomic field since despite being able to identified a large range of amino acid sequence and protein-protein interactions, the functions of individuals protein remain largely unknown. This could revolutionize how research is done, and how many industries can move forward.
With that, I hope that i am able to communicate how this algorithm works, and some future development i am hoping to see.
:)
Slides: [ Ссылка ]
The protein folding problem taken from:
Dill, K. A., Ozkan, S. B., Shell, M. S., & Weikl, T. R. (2008). The protein folding problem. Annual review of biophysics, 37, 289–316. [ Ссылка ]
Image taken from:
1. Graph Mining
[ Ссылка ]
2.Transformer illustration
[ Ссылка ]
3.Levinthal Paradox
[ Ссылка ]
4. AlphaFold 2 Structure
[ Ссылка ]
5. Amino Acid Template
[ Ссылка ]
Thumbnail:
AzaToth(2017)Myoglobin.png@ [ Ссылка ]
Alexbateman(2011)A multiple sequence alignment of the WPP domain.@[ Ссылка ]
Email: liquidbrain.r@gmail.com
Github: [ Ссылка ]
Twitter: [ Ссылка ]